COMPUTATIONAL MODEL FOR SYSTEMATIC BRAIN MAPPING OF MARMOSE
Tracer injection
- The project plans to cover 400 injection sites, one anterograde and one retrograde tracer at each site, evenly distributed across the gray matter of the right hemisphere of the marmoset brain. The stereotaxic coordinates of all injection sites are systematically chosen using an MRI-based atlas (Hashikawa, Nakatomi, & Iriki, 2015).
Cryo-sectioning
- Cryo-sectioning was performed using a Leica CM3050 S Cryostat in a humidity chamber set at 18°C and 64% humidity. This project adapted the modified tape-transfer method (TTM) (Vadim Pinskiy et al., 2015). The slides were transferred to be UV cured for 12 seconds at a UV-LED station setup within the cryostat.
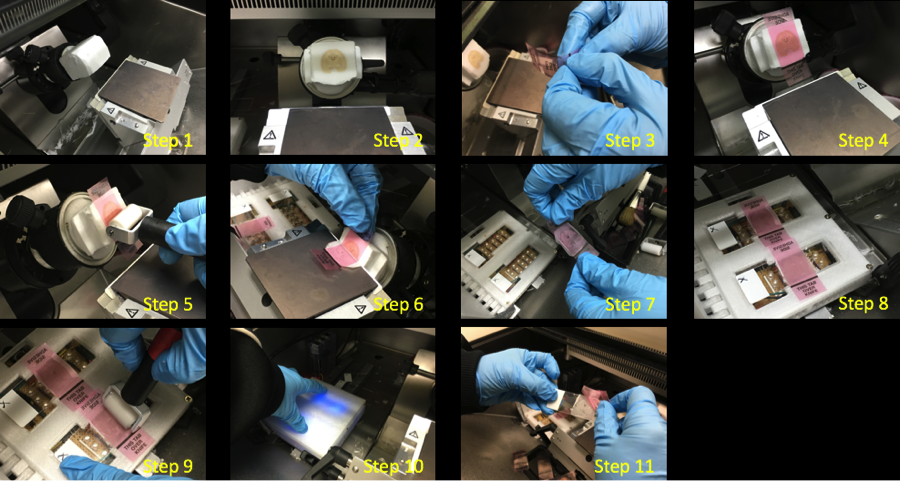

Computational Processing
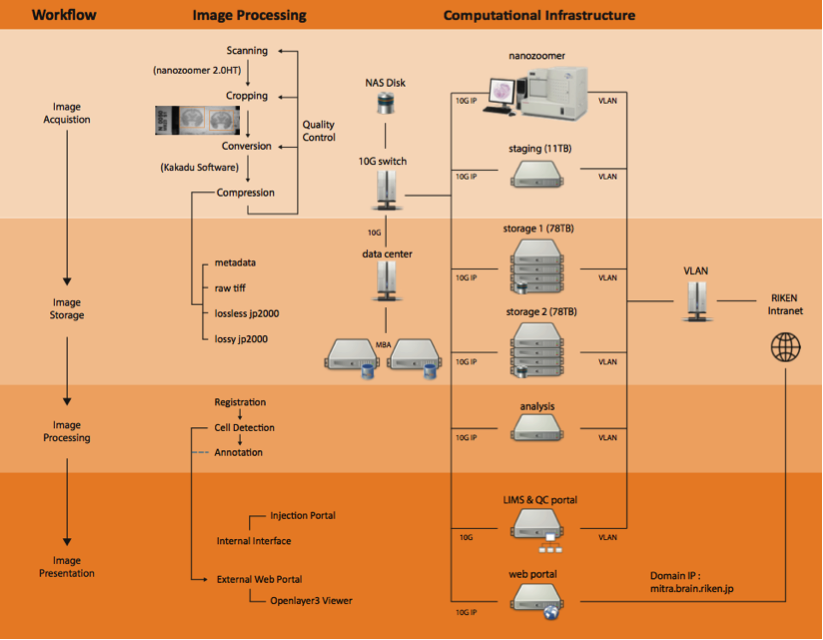
- Computational pipeline with network structure to perform a high-throughput data flow and process. There were four steps of workflow involved in this pipeline including image acquisition, storage, processing and presentation. With this setup, generating a whole marmoset brain dataset with expected high production rate and superior system performance for large data communication became possible.
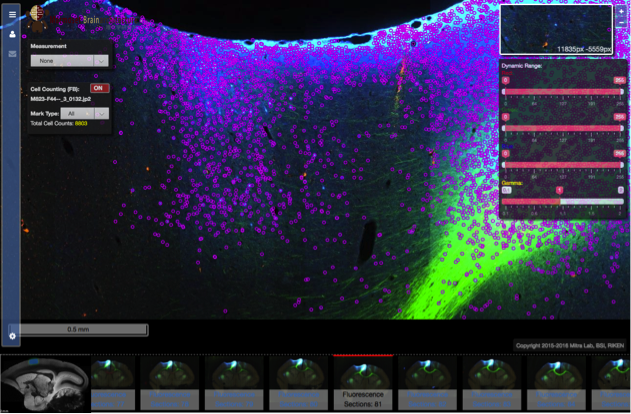
- A web portal had been developed to allow for viewing and interpreting high-resolution JPEG2000 images. The web viewer used an openlayer 3.0 image server and a custom image viewer to allow for fully interactive zoom and pan, supported online adjustment of RGB dynamic ranges, as well measurement and auto cell detection tool.
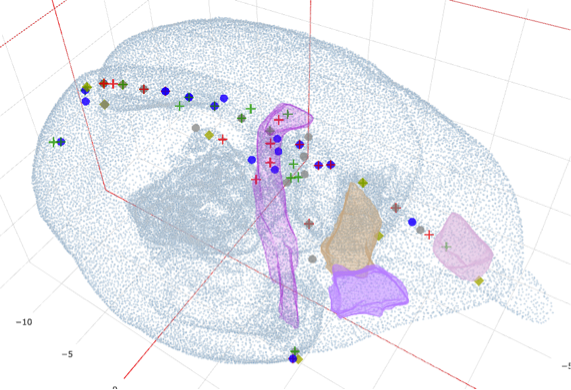
- Automatic annotation of the brain structures was achieved by registering the Hashikawa atlas to post-mortem MRI images using Nissl-Guided Registration with LDDMM and then aligned to the 2D histological brain series. The resulting dataset of the now transformed Hashikawa atlas was subsequently up-scaled, section-by-section along the coronal plane to a high resolution JP2000 file format. The transformed atlas labels were added to the web viewer such that the annotated labels were aligned symmetrically – overlaying the histological image.
[DataID: 2681. Creator Lin Meng Kuan]
